This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Specific Aim Conclusions

The symptoms of schizophrenia can be very disruptive to daily life. While there are treatment methods, including medication and therapy, antipsychotic drugs don't always work for everyone and can also have negative side effects that make their effectiveness in improving quality of life questionable for some schizophrenics. The primary goal of the specific aims presented on this website are to better understand how TOP3B actually leads to schizophrenia, mechanistically. The hope is that this newfound understanding could generate more effective treatment methods of schizophrenia.
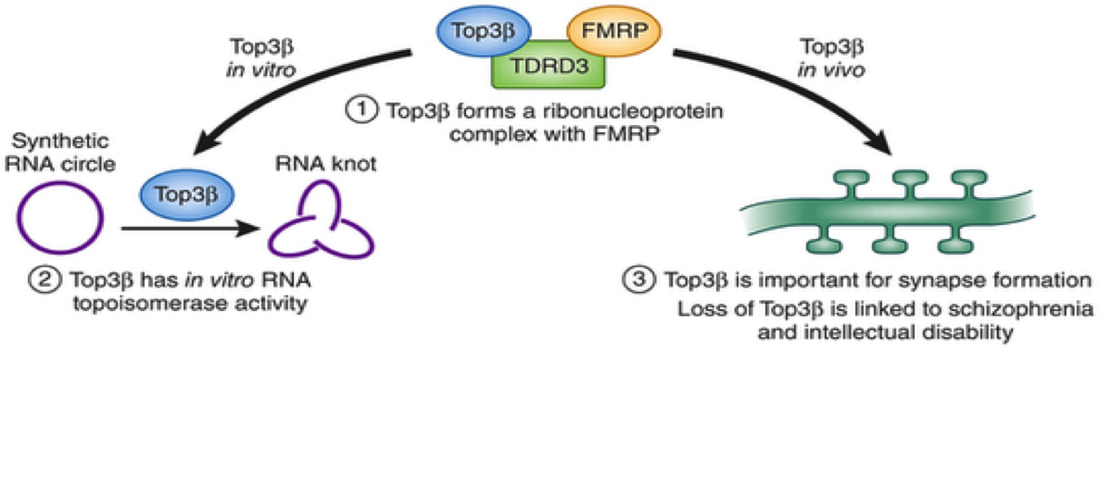
There are three major findings about TOP3B that were used to inform my specific aims. One, that TOP3B has recently been found to bind both other proteins and RNAs [1]. Two, that TOP3β localizes to RNA stress granules, which are aggregates of RNA that form when the cell is under stress [1]. Three, that mice with TOP3B deletions have fewer presynaptic vesicles and fewer synaptic connections that wildtype mice, indicating the TOP3B have a role in neurodevelopment [1]. From these previous findings, I hypothesized that TOP3β would interact with proteins and RNAs involved in stress pathways and that when TOP3B is mutated, these pathways would be interrupted.
 An outline of the methodology for this Aim (Click to Enlarge)
An outline of the methodology for this Aim (Click to Enlarge)
SPECIFIC AIM #1
First, I wanted to compare the genetic sequences of organisms varying from complex to simple nervous systems, I chose Homo sapiens, C. elegans, A. thaliana, and O. sativa. My goal here was to determine genetic function(s) that are important enough to be conserved evolutionarily, which may prove to be important to the development of schizophrenia symptoms. I hypothesized that I would see functions related to stress pathways.
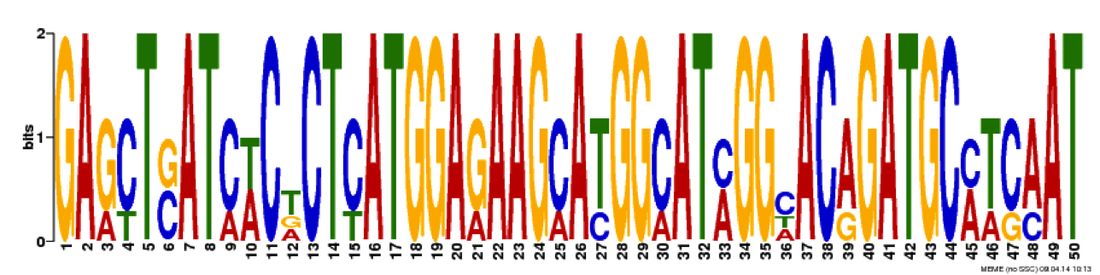
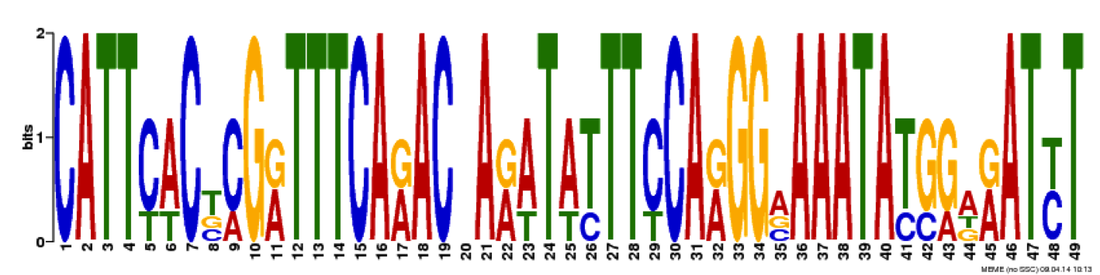
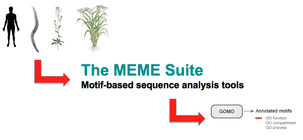
In the image to the right is an outline of what I did for this aim. I took the FASTA sequences of four TOP3B gene homologs and entered them into MEME, which compared the four sequences and finds common motifs. I obtained three motifs, which I then entered into GOMO, which analyzed the motifs based on gene ontology. The three motifs and their GO results can be seen below. I did see some relation to stress pathways in the first motif with one of its biological processes being inflammatory response. However, I also saw a lot of signaling activity across all three motifs (interleuken-2 is a specific cytokine, which is a signaling molecule). TOP3B having an involvement in signaling is logical, given the speech and thought symptoms that can accompany schizophrenia. I carried this observation with me as I worked on my next two aims. I also saw a few surprising results. One, an involvement in the sensory perception of smell in motif one and three. I thought that this could perhaps be taken as a special kind of signaling, as the information must travel from the nose to the brain. Two, an involvement in bacterium response. This was a bit trickier to classify and the best I could do with my hypothesis in mind was this being a response to stress, as bacteria are foreign invaders. Both are certainly points for further investigation.
First, I wanted to compare the genetic sequences of organisms varying from complex to simple nervous systems, I chose Homo sapiens, C. elegans, A. thaliana, and O. sativa. My goal here was to determine genetic function(s) that are important enough to be conserved evolutionarily, which may prove to be important to the development of schizophrenia symptoms. I hypothesized that I would see functions related to stress pathways.
In the image to the right is an outline of what I did for this aim. I took the FASTA sequences of four TOP3B gene homologs and entered them into MEME, which compared the four sequences and finds common motifs. I obtained three motifs, which I then entered into GOMO, which analyzed the motifs based on gene ontology. The three motifs and their GO results can be seen below. I did see some relation to stress pathways in the first motif with one of its biological processes being inflammatory response. However, I also saw a lot of signaling activity across all three motifs (interleuken-2 is a specific cytokine, which is a signaling molecule). TOP3B having an involvement in signaling is logical, given the speech and thought symptoms that can accompany schizophrenia. I carried this observation with me as I worked on my next two aims. I also saw a few surprising results. One, an involvement in the sensory perception of smell in motif one and three. I thought that this could perhaps be taken as a special kind of signaling, as the information must travel from the nose to the brain. Two, an involvement in bacterium response. This was a bit trickier to classify and the best I could do with my hypothesis in mind was this being a response to stress, as bacteria are foreign invaders. Both are certainly points for further investigation.
|
BP: G-protein coupled receptor protein signaling pathway
Regulation of cytokine production Immune Response Inflammatory response SURPRISE: Sensory perception of smell |
BP: Positive regulation of interleukin-2 biosynthetic process
SURPRISE: Bacterium response |
BP: G-protein coupled receptor protein signaling pathway
SURPRISE: Sensory perception of smell |
 An outline of the methodology of this Aim (Click to Enlarge)
An outline of the methodology of this Aim (Click to Enlarge)
SPECIFIC AIM #2
Next, I wanted to identity novel protein interaction of TOP3β in hopes of discovering pathways that TOP3B may influence during the development of schizophrenia. Based on my hypothesis and findings from aim #1, I would expect to see proteins involved in stress and signaling pathways. Here, I chose C. elegans as my model organism because not only is it easy to visual and perform these methods on, but it's simple nervous system can be compared to that of humans.
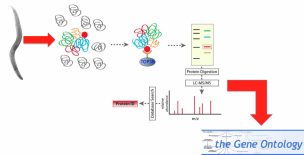
To the right is an outline of the laboratory methods I would use to conduct this aim. With C. elegans, I would perform Tandem Affinity Purification (TAP) followed my Mass Spectrometry (MS) to identify protein binding partners of TOP3β. For a description of TAP-tagging, click here. I would then analyze the recovered proteins based on gene ontology for two reasons. One, to determine the function of these proteins and thus the pathways in which they are involved. Two, to determine if any localize to the brain, which could signify ones of greater importance to schizophrenia development.
Next, I wanted to identity novel protein interaction of TOP3β in hopes of discovering pathways that TOP3B may influence during the development of schizophrenia. Based on my hypothesis and findings from aim #1, I would expect to see proteins involved in stress and signaling pathways. Here, I chose C. elegans as my model organism because not only is it easy to visual and perform these methods on, but it's simple nervous system can be compared to that of humans.
To the right is an outline of the laboratory methods I would use to conduct this aim. With C. elegans, I would perform Tandem Affinity Purification (TAP) followed my Mass Spectrometry (MS) to identify protein binding partners of TOP3β. For a description of TAP-tagging, click here. I would then analyze the recovered proteins based on gene ontology for two reasons. One, to determine the function of these proteins and thus the pathways in which they are involved. Two, to determine if any localize to the brain, which could signify ones of greater importance to schizophrenia development.
|
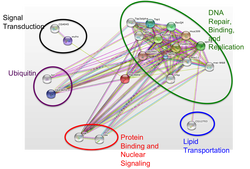
To the left is an STRING diagram that shows the currently known primary and secondary protein interactions of TOP3β. The green section makes sense based on the main function of topoisomerases. The black, red, and purple section are where I would expect to see new interactions from my research methods, as they relate to signaling and perhaps stress (as ubiquitination is often used to mark RNA and proteins for degradation, which could be a response to stress). Scroll down to my future directions section to see more on the lipid transportation (blue) section of this diagram.
|
 An outline of the methodology of the first part of this Aim (Click to Enlarge)
An outline of the methodology of the first part of this Aim (Click to Enlarge)
SPECIFIC AIM #3
For my final aim, I wanted to determine if mutating TOP3B would affect the pathways recovered from aim #2. Furthermore, I wanted to determine if this interruption leads to schizophrenia-like phenotypes, as I predict it would. Once again, I chose C. elegans as my model organism because RNAi can simply be feed to them through E. coli. Also, I found no RNAi results for C. elegans on Wormbase, which makes it an area that needs further study.
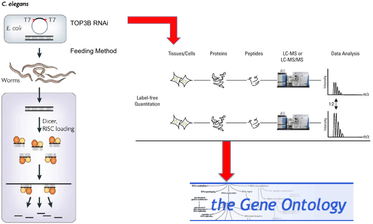
To the right is an outline of my methodology for this aim. I would perform TOP3B RNAi on C. elegans to simulate a TOP3B mutation/deletion. I would then perform quantitative MS on RNAi C. elegans and wildtype C. elegans to compare protein level changes between the two. Again, I would then analyze the proteins with altered levels in the RNAi C. elegans based on GO to determine their function and localization, as they may not be the same as the proteins obtained in the second aim. I think this aim will provide great insight on the mechanism behind how TOP3B leads to schizophrenia symptoms.
For my final aim, I wanted to determine if mutating TOP3B would affect the pathways recovered from aim #2. Furthermore, I wanted to determine if this interruption leads to schizophrenia-like phenotypes, as I predict it would. Once again, I chose C. elegans as my model organism because RNAi can simply be feed to them through E. coli. Also, I found no RNAi results for C. elegans on Wormbase, which makes it an area that needs further study.
To the right is an outline of my methodology for this aim. I would perform TOP3B RNAi on C. elegans to simulate a TOP3B mutation/deletion. I would then perform quantitative MS on RNAi C. elegans and wildtype C. elegans to compare protein level changes between the two. Again, I would then analyze the proteins with altered levels in the RNAi C. elegans based on GO to determine their function and localization, as they may not be the same as the proteins obtained in the second aim. I think this aim will provide great insight on the mechanism behind how TOP3B leads to schizophrenia symptoms.
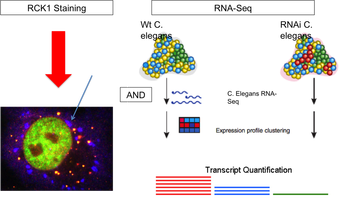 The methodology of the second part of this Aim (Click to Enlarge)
The methodology of the second part of this Aim (Click to Enlarge)
The second part of this aim is measuring the schizophrenia-like phenotype in TOP3B RNAi C. elegans. I would measure this phenotype by analyzing RNA stress granules, which relate both to schizophrenia and recent TOP3B findings. First, I would stain wildtype and RNAi C. elegans for RCK1, a helicase found in RNA stress granules [4]. I would then count the number of granules in each and compare between the two groups. Second, I would perform RNA-Seq on the two groups to quantify the RNA levels and determine if the quantity of RNA known to form stress granules is altered by TOP3B RNAi.
Future Directions
 PolyUnsaturated Fatty Acids
PolyUnsaturated Fatty Acids
The Hashimoto lab recently linked malnutrition and famine during pregnancy to an increased risk of schizophrenia for the developing fetus. As such, they proposed polyunsaturated fatty acids (PUFA) as a potential method of prevention and treatment for the disease [5]. I think this would be a very interesting area for further research. Furthermore, given the STRING finding that TOP3β interacts with a protein involved in lipid transportation, I would like to study not only the effect of PUFA on schizophrenia but on TOP3B. Not only would this provide insight on a potential new wonder drug for this disease, but it could also shed light on the mechanism behind how TOP3B mutation leads to schizophrenia.
| final_presentation.pdf | |
| File Size: | 3300 kb |
| File Type: | |
References
[TOP3B Binding Image] Nott, A. The TOP3B way to untangle RNA. Nat. Neurosci. 2013 Sep: 16(9):1163-64.
[RNA Stress Granules Image] http://www.zmbh.uni-heidelberg.de/stoecklin/
[Synapses Image] http://www.josh-walker.com/2012/12/the-internet-consciousness/
[human] http://imgarcade.com/1/human-being-clipart/
[worm] http://www.k-state.edu/hermanlab/
[arabadopsis] http://www.humanflowerproject.com/index.php/weblog/2006/05/P16
[rice] http://www.eoearth.org/view/article/155736/
[TAP] http://testweb.science.uu.nl/developmentalbiology/interaction_mapping.html
[RNAi] http://www.nature.com/nrg/journal/v9/n7/fig_tab/nrg2364_F1.html
[MS] http://www.piercenet.com/method/quantitative-proteomics
[stress granules] http://www.zmbh.uni-heidelberg.de/stoecklin/
[RNA seq] Sandberg, R. Entering the era of single-cell transcriptomics inbiology and medicine. January 2014. Nature Methods. VOL.11 NO.1
[transcript quantification] http://www.seqwright.com/researchservices/RNA-Seq.html
[1] Nott, A. The TOP3B way to untangle RNA. Nat. Neurosci. 2013 Sep: 16(9):1163-64.
[2] MEME: Multiple Em for Motif Elicitation. Retrieved Apr 15, 2014. http://nbcr-222.ucsd.edu/opal-jobs/appMEME_4.9.113970635652351481237017/meme.html#motif_1
[3] GOMO: Gene Ontology for Motifs. Retrieved Apr 15, 2014. http://nbcr-222.ucsd.edu/opal-jobs/appGOMO_4.9.113970642958481122895160/gomo.html#motif_2
[4] Anderson, P. and Kedersha, N. Stress Granules. 2009. Current Biology. Cell Press. Vol. 19 No. 10. Pp.R397-R398. http://anderson.bwh.harvard.edu/03-Lab%20Publications/03-pdf%20publication%20links/19_Curr_Biol09-May.pdf
[5] Hashimoto, M. et. al. Possibility of polyunsaturated fatty acids for the prevention and treatment of neuropsychiatric illnesses. J Pharmacol Sci. 2014 Mar 18; 124(3):294-300. Epub 2014 Feb 22. PMID: 24561447
[RNA Stress Granules Image] http://www.zmbh.uni-heidelberg.de/stoecklin/
[Synapses Image] http://www.josh-walker.com/2012/12/the-internet-consciousness/
[human] http://imgarcade.com/1/human-being-clipart/
[worm] http://www.k-state.edu/hermanlab/
[arabadopsis] http://www.humanflowerproject.com/index.php/weblog/2006/05/P16
[rice] http://www.eoearth.org/view/article/155736/
[TAP] http://testweb.science.uu.nl/developmentalbiology/interaction_mapping.html
[RNAi] http://www.nature.com/nrg/journal/v9/n7/fig_tab/nrg2364_F1.html
[MS] http://www.piercenet.com/method/quantitative-proteomics
[stress granules] http://www.zmbh.uni-heidelberg.de/stoecklin/
[RNA seq] Sandberg, R. Entering the era of single-cell transcriptomics inbiology and medicine. January 2014. Nature Methods. VOL.11 NO.1
[transcript quantification] http://www.seqwright.com/researchservices/RNA-Seq.html
[1] Nott, A. The TOP3B way to untangle RNA. Nat. Neurosci. 2013 Sep: 16(9):1163-64.
[2] MEME: Multiple Em for Motif Elicitation. Retrieved Apr 15, 2014. http://nbcr-222.ucsd.edu/opal-jobs/appMEME_4.9.113970635652351481237017/meme.html#motif_1
[3] GOMO: Gene Ontology for Motifs. Retrieved Apr 15, 2014. http://nbcr-222.ucsd.edu/opal-jobs/appGOMO_4.9.113970642958481122895160/gomo.html#motif_2
[4] Anderson, P. and Kedersha, N. Stress Granules. 2009. Current Biology. Cell Press. Vol. 19 No. 10. Pp.R397-R398. http://anderson.bwh.harvard.edu/03-Lab%20Publications/03-pdf%20publication%20links/19_Curr_Biol09-May.pdf
[5] Hashimoto, M. et. al. Possibility of polyunsaturated fatty acids for the prevention and treatment of neuropsychiatric illnesses. J Pharmacol Sci. 2014 Mar 18; 124(3):294-300. Epub 2014 Feb 22. PMID: 24561447