This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Protein Homology and Its Importance
|
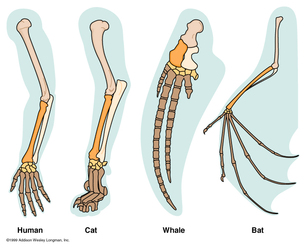
If two things are homologous, it means they share a common ancestor. Protein homology thus refers to proteins that are found in different species and originated from a common ancestor [1]. The term homology was originally coined in the 19th century by Robert Owen, a British comparative anatomist. He used this general term to refer to the similarities between certain structures in different organisms, such as the forelimbs or various vertebrates. It was Charles Darwin who later suggested these similarities were due to a common ancestor [2].
|
The information gained through the study of homology can be extremely useful as it helps us understand how species are related. Homology can be used to help create phylogenetic or evolutionary trees (for more on this see the phylogeny page) [1]. Discovering homologous proteins in simpler organisms also enables us to study these proteins in model organisms, such as mice or flies, to attempt to determine how they function in humans. This can be crucial for disease discovery and treatment.
The Protein Homologs of TOP3β
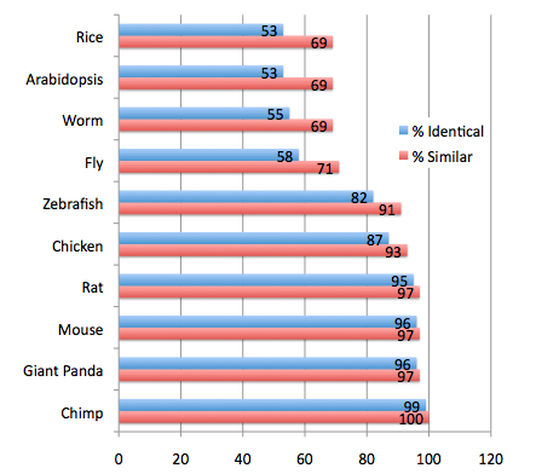
The following protein homologs for the human TOP3β protein were found using BLAST (Basic Local Alignment Search Tool), an online database of gene and protein sequences that uses a sophisticated algorithm to compare sequences between organisms. I simply entered the accession code for the human protein into a protein blast and selected the organisms that I wanted to search for. BLAST then provided me with a list of homologous protein in those organisms, along with two percentages. 'Percent Identical' refers to the extent to which the two sequences have the same amino acids in the same place. 'Percent Similarity' refers to the extent to which two protein sequences are related [3]. The database also presented an Expect (E) value for each percentage; an E value closer to zero means that that number is significant [1].
Analyzing the TOP3β Homologs
The order of most similar to least similar protein as compared to the human homolog is once again, as with the genetic homologs, very logical. This is a highly essential, and thus highly conserved, protein to the basic genetic functions of living organisms. As the chimpanzee is our closest evolutionary relative on this list of organisms, it is to be expected that the proteins would be as similar as they are. The extreme similarity between the chicken, rat, mouse, zebrafish, and giant panda proteins with the human protein is also to be expected as they are all vertebrates. As the fly and worm are invertebrates, it is also logical that their homolog would be less identical to the human homolog than the vertebrate organisms. Yet, the fly and worm proteins are still highly similar, which is extremely useful for research purposes. The same can be said for the two plant species, arabidopsis and rice. Again, the relatively high level of similarity that remains to be seen in these final two species also reiterates the importance of this protein to life.
Below is a list of TOP3β homologs as well as related information. The FASTA link will take you to a page with the protein sequence of that homolog, which was obtained from the protein database of NCBI.
Below is a list of TOP3β homologs as well as related information. The FASTA link will take you to a page with the protein sequence of that homolog, which was obtained from the protein database of NCBI.
|
1. Mus musculus (mouse)-TOP3β
Length: 862 96% identical 97% similar 0% gaps 0.0 E Accession code: EDK97430.1 FASTA 2. Rattus Norvegicus (rat)-TOP3β -1 Length: 862 95% Identical 97% similar 0% gaps 0.0 E Accession code: NP_001099331.1 FASTA 3. Gallus gallus (chicken)-TOP3β -1 Length: 862 87% identical 93% similar 0% gaps 0.0 E Access code: NP_001006181.1 FASTA 4. Drosophila melanogaster (fly)-TOP3β Length: 875 58% identical 71% similar 0% gaps 0.0 E Accession code: NP_511059.2 FASTA 5. Caenorhabditis elegans (worm)-Protein Y48C3A.14 Length: 862 55% identical 69% similar 2% gaps 0.0 E Accession code: NP_001254381.1 FASTA |
6. Danio rerio (zebrafish)-TOP3β -1
Length: 862 82% identical 91% similar 0% gaps 0.0E Accession code: XP_003198651.1 FASTA 7. Pan Trogolodyes (chimp)-TOP3β -1 Length: 862 99% idendical 100% similar 0% gaps 0.0E Accession code: XP_515006.3 FASTA 8. Ailuropoda melanoleuca (Giant panda)-TOP3β -1-like Length: 862 96% identical 97% similar 0% gaps 0.0 E Accession code: XP_002931153.1 FASTA 9. Arabidopsis Thaliana (plant)-DNA topoisomerase, type IA, core Length: 865 53% identical 69% similar 3% gaps 0.0 E Accession code: NP_180760.2 FASTA 10. Oryza Sativa (rice)-Os09g0500600 Length: 866 53% identical 69% similar 3% gaps 0.0 E Accession code: NP_001063578.1 FASTA |
References
[1] Delsuc, F., Brinkmann, H., Philippe, H. Phylogenomic and the reconstruction of the tree of life. Nat Rev Genet. 2005 May; 6(5):361-75.
[2] National Center for Science Education. What is Homology? Created Oct. 2008. Retrieved Feb. 2014. http://ncse.com/evolution/science/what-is-homology
[3] Jan Fassler, Ph.D. and Peter Cooper, Ph.D. NCBI. BLAST Glossary. Created July, 2011. Retrieved Feb. 2014. http://www.ncbi.nlm.nih.gov/books/NBK62051/
[2] National Center for Science Education. What is Homology? Created Oct. 2008. Retrieved Feb. 2014. http://ncse.com/evolution/science/what-is-homology
[3] Jan Fassler, Ph.D. and Peter Cooper, Ph.D. NCBI. BLAST Glossary. Created July, 2011. Retrieved Feb. 2014. http://www.ncbi.nlm.nih.gov/books/NBK62051/