This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
RNA Interference
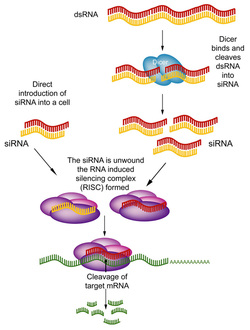 A diagram of RNAi
A diagram of RNAi
RNA interference (RNAi) is an experimental method in which small RNAs bind to mRNAs at a complementary sequence and lead to the degradation of these mRNAs [1]. The process of how these small RNAs are made and used to degrade mRNAs is shown in the diagram to the left. While these small RNAs can be produced endogenously, these small RNAs can also be injected into the cell for experimental purposes. RNAi is a commonly used method to knock down (or greatly decrease) the expression of target gene [1]. The theory behind this methodology is that we can observe the phenotypic results after silencing the expression of this gene in order to determine what this gene's function is. The two major limitations of this method are that it does not 100% knock out the gene (as there are generally some mRNA that remain undegraded) and there can be off target effects. This means that the small RNA could bind to a sequence that is similar to the target sequence and affect the expression of an undesired gene. However, RNAi offers a simple and fast method of gene silencing [1].
RNAi experiments are performed in numerous model organisms. Likewise, there are RNAi databases for each of these model organisms online. Below are the results I obtained when I searched two of these databases for TOP3B RNAi.
RNAi experiments are performed in numerous model organisms. Likewise, there are RNAi databases for each of these model organisms online. Below are the results I obtained when I searched two of these databases for TOP3B RNAi.
RNAi and TOP3B
RNAi for TOP3B could not be found in many of the model organisms. Perhaps this is because TOP3B's connection to schizophrenia is a very recent one. I was able to obtain phenotypic results for TOP3B RNAi in two model organisms: the mouse [2] and fly [3]. The databases I used were MGI (Mouse Genome Informatics) and Flybase.
Analyzing These Findings
Given TOP3B's association with schizophrenia and cognitive impairments, the nervous system defects found in the fly are not surprising. I think that the RNAi results found in the mice are much more interesting. The silencing of TOP3B produced full body inflammation. This could provide insight into how TOP3B functions besides binding DNA. These results may indicate that TOP3B has a role in signaling pathways or immune responses that cause inflammation.
References
[1]. Sugimoto A. (2004). High-throughput RNAi in Caenorhabditis elegans: genome-wide screens and functional genomics. Differentiation. 2004 Mar;72(2-3):81-91. Review.
[2]. MGI. top3b. retrieved Apr 15, 2014. http://www.informatics.jax.org/allele/key/8861
[3]. Flybase. retrieved Apr 15, 2014. http://flybase.org/cgi-bin/cvreport.html?id=FBcv:0000435
[RNAi image] http://www.landesbioscience.com/curie/chapter/4738/
[2]. MGI. top3b. retrieved Apr 15, 2014. http://www.informatics.jax.org/allele/key/8861
[3]. Flybase. retrieved Apr 15, 2014. http://flybase.org/cgi-bin/cvreport.html?id=FBcv:0000435
[RNAi image] http://www.landesbioscience.com/curie/chapter/4738/


